In a comment by Mankiw at Harvard he notes the obvious absurdity of the new Health Care plan. It was written to cover everything. He clearly notes that all other insurance plans cover big items which can be catastrophic. Health Care should be that was as we had argued in our Book a few years ago.
Our recommendations were simply:
1. Universal Coverage: Others should not have to pay for your problems, especially catastrophic ones. Thus we pool our big risks.
2. Individual Coverage: That means like auto and home insurance you personally foot the bill.
3. Mandated Maximum Risk Coverage: This means that if you want to cover everything you can at your own costs but you are covered for the really big things.
4. Payment Support Coverage: For those who cannot afford coverage then the Government may pitch in.
5. Lifestyle Risk Fees: If you chose to smoke, eat in excess, etc then you get to pay more.
6. No "Somatic" Risk Charge: If you were dealt bad genes then you do not get penalized.
Simple, we worked out all the details, it costs much less, and people are happy. Would Washington buy this? No way, they could not control it. Perhaps Mankiw could follow up on this thread, his observation is spot on.
Thursday, March 28, 2013
Wednesday, March 27, 2013
Does This Make Any Sense?
In a recent posting a Health Care promoter was quoted as saying:
"Over the first two years, costs are going to be higher and (the ACA) is going to be seen as a failure” by some, Paduda said. “Of course costs are going to be higher, because of pent-up demand. … But after that, demand (for many) procedures will come down significantly.”
Pent up demand? The person was waiting for three years to have their lung tumor resected, or was it the glioblastoma, that nasty looking melanoma, that renal failure, what pent up demand. Perhaps the rhinoplasty, the botox shots, the extra birth control pills?
"Over the first two years, costs are going to be higher and (the ACA) is going to be seen as a failure” by some, Paduda said. “Of course costs are going to be higher, because of pent-up demand. … But after that, demand (for many) procedures will come down significantly.”
Pent up demand? The person was waiting for three years to have their lung tumor resected, or was it the glioblastoma, that nasty looking melanoma, that renal failure, what pent up demand. Perhaps the rhinoplasty, the botox shots, the extra birth control pills?
In my opinion this makes no sense at all, but it will become the pitch lines to justify additional cost over runs.
The article further states:
“We're focused like a laser on affordability,” American's Health
Insurance Plans CEO Karen Ignagni told Modern Healthcare in an
interview. AHIP has been arguing for months that premiums will rise
because of a new premium tax, essential health benefits required of
individual and small-group plans, and a narrower age-band ratio for
rates. The new ratio means that an older individual will pay no more
than three times more in premiums for the same plan. As a result,
younger people will end up paying more for coverage in 2014 than today.
Now, most states have a 5:1 age-band ratio.
Some of Ignagni's concerns were highlighted in a study released Tuesday by the Society of Actuaries. The actuaries found that the cost of medical claims, one of the key factors driving insurance premiums, will rise an average of 32% nationwide by 2017 as a result of an expected change in the individual market composition. The study further predicts that as many as 43 states could see claims costs increase by percentages in the double digits.
Some of Ignagni's concerns were highlighted in a study released Tuesday by the Society of Actuaries. The actuaries found that the cost of medical claims, one of the key factors driving insurance premiums, will rise an average of 32% nationwide by 2017 as a result of an expected change in the individual market composition. The study further predicts that as many as 43 states could see claims costs increase by percentages in the double digits.
Premiums will rise because providers are mandated to provide a heap of new services and at no cost. It is not just birth control pills but it is every other service known to man and woman. We are looking at in excess of 10% pa compounded rates of increase. There is no real incentive to drive down costs in such areas as obesity driven Diabetes. In fact the law accommodates that and pays for any related costs.
As we had indicated when this mess began, we expect explosive cost growth going right to the National Debt!
Labels:
Health Care
The Incompetence of the Health Care Insurers
First rule of life is: Do Not Get Sick, Ever, Just Die, Neatly.
Otherwise you have to deal with health care providers and insurers. Today I tried to reorder some eye drops. Not real real critical, but timely. So I went off to the website of the insurer to do so;
1. Ten days ago I ordered on line and it seemed simple. Just renew it.
2. Yesterday I found that they cancelled the prescription order.
3. Then I emailed back and forth to see what was wrong but to no avail. First we all know email really does not work in communicating, and second their objective is not to honor the agreement. So obfuscate.
4. Today after 10 min with an electronic operator saying names and numbers a half a dozen time I go through to a human. Apparently they believe that I can only get so much a year no matter what.
5. The problem, about six months ago they switched from an American supplier to an Indian supplier. The Indian applicator leaks drops all over so one goes through the therapeutic at twice the rate. But the drug is most likely half the price, but if you use it at twice the rate... Do not expect some moron at an insurer to recognize that 0.5 time 2 is still one!
6. The putative solution is to double the dosing, namely have the physician write a script for qid not bid and then hope their computer honors it. Do not count on it.
7. The alternative is to go over into Canada and buy it there. Despite the rumors they really do not ration, that much.
Well this is another sign that we will be seeing a total disaster in Health Care.
Otherwise you have to deal with health care providers and insurers. Today I tried to reorder some eye drops. Not real real critical, but timely. So I went off to the website of the insurer to do so;
1. Ten days ago I ordered on line and it seemed simple. Just renew it.
2. Yesterday I found that they cancelled the prescription order.
3. Then I emailed back and forth to see what was wrong but to no avail. First we all know email really does not work in communicating, and second their objective is not to honor the agreement. So obfuscate.
4. Today after 10 min with an electronic operator saying names and numbers a half a dozen time I go through to a human. Apparently they believe that I can only get so much a year no matter what.
5. The problem, about six months ago they switched from an American supplier to an Indian supplier. The Indian applicator leaks drops all over so one goes through the therapeutic at twice the rate. But the drug is most likely half the price, but if you use it at twice the rate... Do not expect some moron at an insurer to recognize that 0.5 time 2 is still one!
6. The putative solution is to double the dosing, namely have the physician write a script for qid not bid and then hope their computer honors it. Do not count on it.
7. The alternative is to go over into Canada and buy it there. Despite the rumors they really do not ration, that much.
Well this is another sign that we will be seeing a total disaster in Health Care.
Labels:
Health Care
Monday, March 25, 2013
Cancer and New Therapeutics
There is an explosion of new cancer therapeutics. About ten years ago we saw imatinib for CML and now we have quite a few for metastatic melanoma, once a terminal disease for certain. In the Melanoma case we see some 20% may survive for extended periods of time. However the average life extension may be only 6 months at a cost that may exceed $100K. In addition there may need be several of these therapeutics used at one time.
A former Administration Health Care adviser has written on this of late:
Many cancer patients, after getting a diagnosis of a terrifying disease, pursue any potentially promising therapy, regardless of the price. But the main cost driver is the fee-for-service payment system. The more doctors do for patients, the more reimbursement they receive. Surgeons earn more for every procedure. Oncologists typically make more money if they use newly approved drugs and the latest radiation treatments than if they use cheaper, older alternatives that work just as well. (This is because they get paid back the cost of the drug, in addition to an extra 6 percent of that cost — the more expensive the drug, the higher the compensation.)
His point is the 6% on the $100K charge. That is $6K per patient per six months. Take melanoma. The incidence is about 75K per year. Of that some 12K to 20K it the drug profile. At say $100K per person and assuming all persons, 20K, we would have in any one year $2B in costs and $120M paid to Oncologists. Is that too much? I guess it depends if you are in the 20% or the 80%.
He suggests changes:
First, over the next few years, the payment system needs to move away from fee-for-service toward a system of bundled payments, in which doctors are paid one fee for all the treatments involved in caring for a cancer patient.
Point well taken. But the problem is the way we compensate people based upon past assumptions.
Second, insurers have to give physicians information about where they are spending money.
I would suggest the patient also be informed. Patients all too often assume that the costs of the medication is low. They have no idea what the costs are. Moreover the basis for the costs should also be known. One must remember that the drug companies have gone through multi-Phase trials of hundreds of patients each at tens of thousands per patients just to the management of the Trial. Recall that the CROs, the Clinical Research Organizations, generate almost $30B in annual revenue just managing the Trials to comply with the FDA. That is not money in the pockets of the Pharmas.
Third, any change in payment methods must be accompanied by rigorous quality monitoring to ensure that there is neither under- nor over-utilization of care.
Quality, now just what do we mean by that? This is what drove the character nuts in the Zen and the Art of Motorcycle Maintenance. Really, is it nothing more than what is in the eye of the beholder.
Fourth, we need more “high touch” oncology practices. In these practices, nurses manage common symptoms before they escalate to the point that they require visits to the emergency room..
Part of this is that the Oncologists are dealing with a mass amounts of new and different genetically targeted drugs which address pathways that they may have never been exposed to in Medical School. One melanoma drug leads to a new skin cancer, an unexpected effect.
Fifth, we need better incentives for research. Many expensive tests and treatments are introduced without evidence that they improve survival or reduce side effects, and with poor information about which patients should receive them.
Here I would disagree. The Trials are somewhat extensive but when you apply something used over 600 people to 20,000 you get a whole new set of issues. A drug may have to be withdrawn.
One key question is who should receive the new therapeutics? How do we manage them? Cancer is terrifying to the patient. But we now have an environment where people can find out about these new medications and demand them. Physicians are then pressed to use them, albeit with little significant survival benefit, on average. Yet that 20% who do survive contain valuable information for the next step.
Thus do we view use of the new therapeutics as the cost of continuing research or the cost of providing care?
A former Administration Health Care adviser has written on this of late:
Many cancer patients, after getting a diagnosis of a terrifying disease, pursue any potentially promising therapy, regardless of the price. But the main cost driver is the fee-for-service payment system. The more doctors do for patients, the more reimbursement they receive. Surgeons earn more for every procedure. Oncologists typically make more money if they use newly approved drugs and the latest radiation treatments than if they use cheaper, older alternatives that work just as well. (This is because they get paid back the cost of the drug, in addition to an extra 6 percent of that cost — the more expensive the drug, the higher the compensation.)
His point is the 6% on the $100K charge. That is $6K per patient per six months. Take melanoma. The incidence is about 75K per year. Of that some 12K to 20K it the drug profile. At say $100K per person and assuming all persons, 20K, we would have in any one year $2B in costs and $120M paid to Oncologists. Is that too much? I guess it depends if you are in the 20% or the 80%.
He suggests changes:
First, over the next few years, the payment system needs to move away from fee-for-service toward a system of bundled payments, in which doctors are paid one fee for all the treatments involved in caring for a cancer patient.
Point well taken. But the problem is the way we compensate people based upon past assumptions.
Second, insurers have to give physicians information about where they are spending money.
I would suggest the patient also be informed. Patients all too often assume that the costs of the medication is low. They have no idea what the costs are. Moreover the basis for the costs should also be known. One must remember that the drug companies have gone through multi-Phase trials of hundreds of patients each at tens of thousands per patients just to the management of the Trial. Recall that the CROs, the Clinical Research Organizations, generate almost $30B in annual revenue just managing the Trials to comply with the FDA. That is not money in the pockets of the Pharmas.
Third, any change in payment methods must be accompanied by rigorous quality monitoring to ensure that there is neither under- nor over-utilization of care.
Quality, now just what do we mean by that? This is what drove the character nuts in the Zen and the Art of Motorcycle Maintenance. Really, is it nothing more than what is in the eye of the beholder.
Fourth, we need more “high touch” oncology practices. In these practices, nurses manage common symptoms before they escalate to the point that they require visits to the emergency room..
Part of this is that the Oncologists are dealing with a mass amounts of new and different genetically targeted drugs which address pathways that they may have never been exposed to in Medical School. One melanoma drug leads to a new skin cancer, an unexpected effect.
Fifth, we need better incentives for research. Many expensive tests and treatments are introduced without evidence that they improve survival or reduce side effects, and with poor information about which patients should receive them.
Here I would disagree. The Trials are somewhat extensive but when you apply something used over 600 people to 20,000 you get a whole new set of issues. A drug may have to be withdrawn.
One key question is who should receive the new therapeutics? How do we manage them? Cancer is terrifying to the patient. But we now have an environment where people can find out about these new medications and demand them. Physicians are then pressed to use them, albeit with little significant survival benefit, on average. Yet that 20% who do survive contain valuable information for the next step.
Thus do we view use of the new therapeutics as the cost of continuing research or the cost of providing care?
Labels:
Cancer
Other Models for Cancer Propagation
As we have described before in our diffusion model and in our Markov chain enhancement, cancer can be modeled as a complex process of; (i) diffusion, (ii) propagation, (iii) proliferation, and (iv) mutation. In each of the above we have defined and measurable processes that result from external and internal physiological and genetic known paths. For example we know that cancer cells when propagated to say the liver will use the liver cells to increase propagation, and in addition will also use liver extracellular factors to potentially suppress gene expression, a factor that appears as a loss of a gene in a pathway.
In a recent paper by Newton et al the authors have developed an alternative ad hoc model, independent of pathways. They state:
The classic view of metastatic cancer progression is that it is a unidirectional process initiated at the primary tumor site, progressing to variably distant metastatic sites in a fairly predictable, though not perfectly understood, fashion.
A Markov chain Monte Carlo mathematical approach can determine a pathway diagram that classifies metastatic tumors as 'spreaders' or 'sponges' and orders the timescales of progression from site to site. In light of recent experimental evidence highlighting the potential significance of self-seeding of primary tumors, we use a Markov chain Monte Carlo (MCMC) approach, based on large autopsy data sets, to quantify the stochastic, systemic, and often multi-directional aspects of cancer progression.
We quantify three types of multi-directional mechanisms of progression: (i) self-seeding of the primary tumor; (ii) re-seeding of the primary tumor from a metastatic site (primary re-seeding); and (iii) re-seeding of metastatic tumors (metastasis re-seeding). The model shows that the combined characteristics of the primary and the first metastatic site to which it spreads largely determine the future pathways and timescales of systemic disease.
For lung cancer, the main `spreaders' of systemic disease are the adrenal gland and kidney, whereas the main `sponges' are regional lymph nodes, liver, and bone. Lung is a significant self-seeder, although it is a `sponge' site with respect to progression characteristics.
Now we believe that this is an ad hoc model because it fails to allow for the inclusion of the cellular pathway dynamics expressly. We believe that it is essential to have the underlying "system model" based on reality as an integral part of the essential model and not just posit an ad hoc model. Our approach has that at its core.
The advantages of including such a model are as follows:
(1) The inclusion allows for the validation based on reality.
(2) The model allows for the inclusion of a model useful for controllability and observability factors. Namely it facilitates the ability to predict usefulness of therapeutics.
(3) A model where the coefficients can be estimated from data and where the estimation is based upon a known physical reality. Thus diffusion is key when say in melanoma we lose E cadherin and thus a melanocyte starts to drift, and where in melanoma we lose RAF control and proliferation begins to occur, or in prostate cancer where in the bone marrow we have excess growth factors and enhanced receptors allowing the malignant prostate cell to proliferate. The nexus with the details of reality are a key and essential factor, a sine qua non, for any such model.
Thus although models of this type may mimic reality I feel that they lack the true essence of reality. One of the essential elements of control theory is the ability to have models of reality which are predicated on essential underlying physical truth.
Labels:
Cancer
Get a Job
There is an article on today's NY Times regarding the value of an engineering education. As usual there are the liberal arts types telling us we should be broadly educated as a citizen first and then be productive second.
Their definition of being educated is being told by them what they believe is correct. One can read Locke by one's self, and one does not need some Marxist Professor telling us of the oppression of the working class by the capitalists. After all it was Locke who identified ownership in property with labor.
One Professor of poetry writes:
You are asking the wrong question. Today’s students should not be chasing “return on investment.” They should be considering a different “R.O.I.,” over the longer term: return on the individual. That means a lifetime of skills, like learning how to learn, adapting to a changing economy and society, expressing oneself clearly, solving problems creatively, and acquiring the civic skills to contribute to our democracy.
Colleges and universities want graduates to thrive in their lives, to see the value of the degree. But we must enlarge and enrich our definition of “value” beyond the purely monetary. Students must be prepared for a lifetime of engagement, not for specific jobs that may change or even disappear. The need to innovate goes deeper than the job market; it reaches into how we organize our social and political interactions.
How in God's name can you "thrive" in your life if you cannot support yourself. And your offspring.
Once you have a good job, a sustainable profession, and hopefully independent, then you can be free to do whatever you want. I read philosophy, literature, history, but these became so much more critical when I lived them in my business life. The Thirty Years War became real when the Swedes tried to crush my business and I was saved by the French, despite the fact that I was in Prague but the business was in Poland! Catholics versus Protestants, the Thirty Years War in real life. My ancient Greek became current Greek, since I had to do deals in Athens.
The critical skills to contribute to our democracy is not spending one's life reading poetry but in contributing first hand to is economic survival, and that is done by producing.
Again I recount my road to MIT was from a trash can working as a part time maintenance worker at a New York City school, finding the MIT catalog, and from there the tale is well known. If I did not have my hands in that trash can emptying out the old catalogs perhaps a different life. Perhaps. It was not the poetry, it was not Yeats, it may have been Homer.
Their definition of being educated is being told by them what they believe is correct. One can read Locke by one's self, and one does not need some Marxist Professor telling us of the oppression of the working class by the capitalists. After all it was Locke who identified ownership in property with labor.
One Professor of poetry writes:
You are asking the wrong question. Today’s students should not be chasing “return on investment.” They should be considering a different “R.O.I.,” over the longer term: return on the individual. That means a lifetime of skills, like learning how to learn, adapting to a changing economy and society, expressing oneself clearly, solving problems creatively, and acquiring the civic skills to contribute to our democracy.
Colleges and universities want graduates to thrive in their lives, to see the value of the degree. But we must enlarge and enrich our definition of “value” beyond the purely monetary. Students must be prepared for a lifetime of engagement, not for specific jobs that may change or even disappear. The need to innovate goes deeper than the job market; it reaches into how we organize our social and political interactions.
How in God's name can you "thrive" in your life if you cannot support yourself. And your offspring.
Once you have a good job, a sustainable profession, and hopefully independent, then you can be free to do whatever you want. I read philosophy, literature, history, but these became so much more critical when I lived them in my business life. The Thirty Years War became real when the Swedes tried to crush my business and I was saved by the French, despite the fact that I was in Prague but the business was in Poland! Catholics versus Protestants, the Thirty Years War in real life. My ancient Greek became current Greek, since I had to do deals in Athens.
The critical skills to contribute to our democracy is not spending one's life reading poetry but in contributing first hand to is economic survival, and that is done by producing.
Again I recount my road to MIT was from a trash can working as a part time maintenance worker at a New York City school, finding the MIT catalog, and from there the tale is well known. If I did not have my hands in that trash can emptying out the old catalogs perhaps a different life. Perhaps. It was not the poetry, it was not Yeats, it may have been Homer.
Labels:
Commentary,
Economy,
Education
Wednesday, March 20, 2013
Melanoma and TVEC
Amgen announced success with Phase 3 Trial using TVEC, Talimogene Laherparepvec, which is an elegant approach to managing melanoma. The simple idea, and most great ideas are simple, is that you use a virus to penetrate specific cells, and then get the virus to do what it does best, multiply, and kill the cell.
Amgen describes TVEC as:
Talimogene laherparepvec is an investigational oncolytic immunotherapy designed to selectively replicate in tumor tissue. Talimogene laherparepvec is injected directly into tumor tissue and then replicates until the membrane of the cancer cells rupture, thereby destroying the cells, in a process known as cell lysis. The virus that was contained in these cells is then released locally in the tumor tissue along with GM-CSF, a white blood cell growth factor that the virus is engineered to express. This is intended to lead to the activation of a systemic immune response to kill tumor cells throughout the body.
As is reported in GEN:
TVEC is based on a herpes simplex virus type 1 (HSV-1) that has been modified to selectively replicate in tumor cells without harming healthy cells. TVEC represents a new class of therapy against melanoma because it combines local with systemic effects against tumors. That effect is believed to be achieved through the functional deletion of two key genes (ICP34.5 and ICP47) from HSV-1, which deprives it of the proteins normally used for circumventing the body’s response to infections, followed by the addition of the human GM-CSF gene.
When injected directly into tumors, TVEC selectively replicates until the membrane of the cancer cells rupture or lyse, destroying these cells and releasing the viruses that have been replicated. The released viruses, in turn, invade more tumor cells—a cycle that continues until the weakened virus encounters healthy cells. During replication, TVEC also induces the tumor to produce GM-CSF—a white blood cell growth factor that attracts and activates the cells required for a systemic immune response.
This elegant approach selects a viral carrier which selects the melanoma cell and then enters it and replicates itself until the cell is killed off. This approach may have great applications in many other cancers.
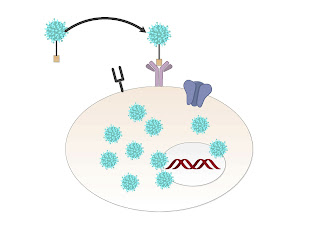
Simply it works as follows:
Step 1 Virus is engineered to have attraction to a melanoma cell.
Amgen describes TVEC as:
Talimogene laherparepvec is an investigational oncolytic immunotherapy designed to selectively replicate in tumor tissue. Talimogene laherparepvec is injected directly into tumor tissue and then replicates until the membrane of the cancer cells rupture, thereby destroying the cells, in a process known as cell lysis. The virus that was contained in these cells is then released locally in the tumor tissue along with GM-CSF, a white blood cell growth factor that the virus is engineered to express. This is intended to lead to the activation of a systemic immune response to kill tumor cells throughout the body.
As is reported in GEN:
TVEC is based on a herpes simplex virus type 1 (HSV-1) that has been modified to selectively replicate in tumor cells without harming healthy cells. TVEC represents a new class of therapy against melanoma because it combines local with systemic effects against tumors. That effect is believed to be achieved through the functional deletion of two key genes (ICP34.5 and ICP47) from HSV-1, which deprives it of the proteins normally used for circumventing the body’s response to infections, followed by the addition of the human GM-CSF gene.
When injected directly into tumors, TVEC selectively replicates until the membrane of the cancer cells rupture or lyse, destroying these cells and releasing the viruses that have been replicated. The released viruses, in turn, invade more tumor cells—a cycle that continues until the weakened virus encounters healthy cells. During replication, TVEC also induces the tumor to produce GM-CSF—a white blood cell growth factor that attracts and activates the cells required for a systemic immune response.
This elegant approach selects a viral carrier which selects the melanoma cell and then enters it and replicates itself until the cell is killed off. This approach may have great applications in many other cancers.
Simply it works as follows:
Step 1 Virus is engineered to have attraction to a melanoma cell.
Step 2 The virus invades.
Step 3 The virus kills the targeted cell.
Nice idea and execution. Very interesting execution and great to follow up on. I tried something akin to this a few years ago on plant cells, TMV vector, with some success.
Labels:
Cancer
Monday, March 18, 2013
Cancer Cell Dynamics
We have previously introduced a cancer cell propagation
model in earlier discussions which others have also considered. However our
model is for a single cell type which grows diffuses and flows. The rates of
each are dependent on where the cell is.
In this brief note we add other elements, namely the
probability that a cell can mutate and that as it mutates the factors related
to the propagation model may also change. We calculate a similar diffusion
equation now for the average number of malignant cells by region and by type.
That is we demonstrate in the following graphic summary:
1. The standard diffusion-flow-proliferation model applies
on a per-region and per cell type basis. This means that the constants we have
developed previously will depend on the specific cell type as well, namely how
many mutations have occurred.
2. That we know there are multiple mutations in cancer
cells. Some may have a few and are indolent and others may have many and be
aggressive. We develop a Markov model for such cell progression.
3. We combine the three element spatio-temporal model with
the Markov cell mutation model and this allows us to determine the average
number of cells of a specific type in any part of the body at any point in
time.
4. We then discuss how one may use this model for prognostic
and therapeutic purposes.
The main observation in this brief section is that the
average number of malignant cells of a specific mutation state can be
determined by the following:
In this equation the n is an NX1 vector of average numbers
in spatio-temporal dependent values of each of N possible mutations and the L
value is the spatio-temporal dependent operator matrix and Λ is a matrix
describing the Markov transition probabilities between mutations.
It should be clear that we can measure all of the constants
involved and thus determine the result. As a counter-distinction we can measure
the n values and mutation states and determine the constants.
The expanded model considers the issue diagrammed below:
The next issue is the ability to determine what the factors
are in the specific model, namely the values of the constants, and secondly the
validation of the model itself.
Thus there are two dimensions of issues here:
1. Model Identification and Validation: In previous work we
referred to this as the Observability problem. Namely if we have a model and we
can identify the required parameters, then can this model be used to determine
the end state which will be attained. This is the prognostic problem.
2. Model Utilization: As with the previous cases, if we have
this model, and we have identified the constants, can we determine actions which
may be taken to control the end state of the system? This is the
Controllability problem. It states that perhaps having such a model we can
determine methods and means to drive the system, in this case the average
number of malignant cells of genotype say G, to a new end state, one where we
have reduced the number of bad cells to a de minimis level. This is the
therapeutic problem.
There also is a third element:
3. Identification: In both of the two previous issues we
assumed that there existed a method by which we could determine the constants
of diffusion et al and furthermore that we could ascertain the list of possible
mutations, and also their Markov transition probabilities. This may be
accomplished in two ways. First, we can accomplish this by in vitro studies.
Second, we can achieve this by using the model itself in a classic system
identification model with in vivo analyses.
Thus the analysis contained herein is an initiation of what
appears to be an innovative way to look at cancer. There have been many studies
in more specific and segmented areas but there has not to my knowledge been a
study that has examined cancer in such a broad and overarching manner. In
essence we have included all of the variables that one may ask for.
To better understand we depict the progression of prostate
cancer below.
Step 1: Benign State, here we have five segments; prostate,
two tissue-blood barriers, blood, and bone.
Step 2: We have the beginning of a cancer due to some
mutation of the basal or luminal cells. The cancer proliferates and diffuses.
It is still localized here.
Step 3: Diffusions sends cancer cells to the blood barrier.
Step 4: The blood barrier is crossed, and we assume by
diffusion. Across this barrier there is no proliferation or flow, just
diffusion.
Step 5: The blood barrier is crossed and the cell is now in
the blood stream. Here we have flow but no diffusion and no proliferation.
Step 6: The blood barrier is crossed again as discussed
above.
Step 7: Metastasis is complete by having the new malignant
cells in the bone and proliferation and diffusion predominate.
The above steps are common is almost all cancers. The
assumptions here are:
1. The same malignant cell moves across the body.
2. Each separate area, in this case five, has constant
diffusion, flow and proliferation constants.
3. That we can then measure the number of cells from this
deterministic model.
In the case where they are uniform constants we can solve
the equation. In the case where they are uniform constants across uniform
spatial domains then we can also solve the equations evoking boundary
conditions.
We now want to expand this model to include multiple
malignant cell types. Also we want to include their stochastic dynamics as
well.
Consider a cell with five possible mutations. We show the
genes below. The call may begin with one mutation and then move to a second and
so forth. Each path is assumed to be possible and the results of each path are
different.
Now we can consider a model for the above simple example. We
have 5 possible mutations and they may occur in any order. We assume they occur
one at a time. We can identify any number of cells as n(x,t), as the number of
cells after one mutation at location x and at time t, of mutation k.
Now we have the following observations:
1. At mutation 1 we have 5 possible cell mutants.
Furthermore each may be considered a cancer cell and the growth, diffusion and
flow are as described above. Some of the mutations may be indolent and some
aggressive.
2. At mutation 2 we have 5*4 possible cells. The question is
that some are say PTEN then cMyc or cMyc then PTEN. Are they the same, and this
means the difference between perturbation and combination? Are they distinct by
have been ordered differently or are they the same? If it is a combination we
have 10 instead of 20 different mutations.
3. At mutation 3 we have 5*4*3 and at 4 we have 5*4*3*2 or
120 permutations.
4. At any location we may have any one or a combination of
these mutation types. There are two factors driving their number:
a. A single type will have growth, dispersion and movement
dynamics with the above mentioned model but each mutation will respond
differently since their coefficients will be different. Some may grow faster
and some may diffuse faster. There is no a priori ranking of the coefficients.
b. The surrounding mutant types will also tend to mitigate
growth.
For example consider the following three gene mutation case:
We can thus make several important observations regarding
this model.
1. Prognostic and Therapeutic: We can determine the
transitions and the factors related to diffusion, flow and growth. Thus we can
use the result as a powerful one for prognostic and therapeutic results. As we
had indicated earlier, the Observability and Controllability issues are
essentially Prognostic and Therapeutic respectively.
2. Variances: The results are for the average. We can
determine the results for the variances as well. We have examined the variances
on the averages and they are somewhat complex and we do not believe that they
lend significant additional information at this time.
3. Solutions: The solutions to these equations are readily
obtained using standard techniques. They can, in addition, be determined in
closed form results.
Labels:
Cancer
Why I Left the IEEE
The IEEE is the professional organization which represents Electrical Engineers. In 1961 I believe I joined the IEEE and remained a member until 2005. At that point I was quite disappointed in the fact that publications had become group affairs and that worse the articles were just repeats of the same idea, but now promulgated by a group. Further the group was usually headed by some senior faculty member who sat on some IEEE committee.
Now in today's NY Times the representative of the IEEE testifies before Congress seeking gender equality in Visas for Electrical Engineers. It states:
Let me provide some facts. At MIT the predominant number of Visa eligible students are men. Did MIT discriminate? Hardly! They were the best ones who applied.
Is there a gender imbalance in Electrical Engineering? Perhaps that is the wrong question. You see EE is hard. Yes really hard, and it is competitive, really competitive. So you get out what people put in. We see many Asian students, especially at the PhD level. Of my last 8 Graduate students only one is left in the US, all were foreign students, and some of the best are now in our competitors, and only one was female. Those are facts. What the IEEE is complaining about seems utter nonsense. The problem is not issuing more Visas to men than women, in fact I would be willing to wager that percentage wise of graduates more women get Visas than men.
Let us examine the logic and consequence of her apparent suggestion, in my opinion.
First, if as I suspect, the males in EE represent well over 70% of PhDs and females 30% or less,
Then, if she demands parity in percents in terms of Visas,
One concludes that women will receive twice as many visas as men, and them men will be sent back from whence they came and produce value there and not here.
Engineering is not law or medical school, it still is male dominated. The choice or selection was made decades before graduation. Each and every one of those graduates should be induced to stay here and be creative, not the social engineering selective process of some person at some "professional" society which represents God knows what.
No wonder I left the IEEE! My dues paid for people like this one.
Now in today's NY Times the representative of the IEEE testifies before Congress seeking gender equality in Visas for Electrical Engineers. It states:
Karen Panetta, the vice president for communications and public
awareness for the Institute of Electrical and Electronics Engineers in
the United States of America, will testify that “the vast majority of
H-1B workers are men,” according to her prepared remarks.
Ms. Panetta’s testimony points to “a serious gender imbalance in
science, technology, engineering and math” as part of the reason that
these H-1B visas in high-tech fields skew disproportionately toward men.
But she also adds, “If a major immigration program effectively
discriminated based on race or national origin, would that be O.K.?”
Let me provide some facts. At MIT the predominant number of Visa eligible students are men. Did MIT discriminate? Hardly! They were the best ones who applied.
Is there a gender imbalance in Electrical Engineering? Perhaps that is the wrong question. You see EE is hard. Yes really hard, and it is competitive, really competitive. So you get out what people put in. We see many Asian students, especially at the PhD level. Of my last 8 Graduate students only one is left in the US, all were foreign students, and some of the best are now in our competitors, and only one was female. Those are facts. What the IEEE is complaining about seems utter nonsense. The problem is not issuing more Visas to men than women, in fact I would be willing to wager that percentage wise of graduates more women get Visas than men.
Let us examine the logic and consequence of her apparent suggestion, in my opinion.
First, if as I suspect, the males in EE represent well over 70% of PhDs and females 30% or less,
Then, if she demands parity in percents in terms of Visas,
One concludes that women will receive twice as many visas as men, and them men will be sent back from whence they came and produce value there and not here.
Engineering is not law or medical school, it still is male dominated. The choice or selection was made decades before graduation. Each and every one of those graduates should be induced to stay here and be creative, not the social engineering selective process of some person at some "professional" society which represents God knows what.
No wonder I left the IEEE! My dues paid for people like this one.
Labels:
Commentary
Saturday, March 16, 2013
Google Reader and How People Think
I have been thinking and reading a bit about the response to the termination of Google Reader and I have been a bit surprised, but should not really have been, by those who felt it was no loss. For them perhaps. But Google Reader is an artifact of how some people think.
You see my computer screen is organized for efficiency. I have folders in the center of the screen, not just piling up as Microsoft suggests and filling up the screen. Mine are alphabetically organized and then within each is specific applications. Files are organized by topics, subtopics, etc. Google Reader is also organized that way.
I want to watch as topics may come by, I could care less about the images, unless I seek out the specific article. Google Reader let's me know what is new say in a specific area and as some reader may not I often then write upon that in detail, for better or worse. I leave that up to the readers.
But sifting through masses of online data real time to see what has become new and how to fit that in to current research and writings is one of my organized ways. Twitter is totally useless. I have an account, I had to get one for something that I now forgot. I have Linkedin, and it has become the Ginza strip, all bright and useless. I left Facebook years ago, and Google+ makes no sense at all. There just seems to be stuff pasted by others and I have no idea why.
But if you are goal directed, with an organized mental bent, then Google Reader was spot on. If you are a Facebook fanatic, then it would make no sense. Google Reader was for those who did something with what they obtained, Facebook is a narcotic. It is Crazy Bird of the social set.
Google Reader led you to new ideas which made you modify and expand old ideas, it led to challenges, it led to thought. Google Reader was for the organized mind, not the idle mind.
Thus there is a potential wealth of information on how different people think. Social media is for those who want others to think for them. Perhaps a great marketing tool. Google Reader, plain and simple, is for those who want to think for themselves. This tells me something about the folks at Google. They are changing, and perhaps not for the best. Pity!
You see my computer screen is organized for efficiency. I have folders in the center of the screen, not just piling up as Microsoft suggests and filling up the screen. Mine are alphabetically organized and then within each is specific applications. Files are organized by topics, subtopics, etc. Google Reader is also organized that way.
I want to watch as topics may come by, I could care less about the images, unless I seek out the specific article. Google Reader let's me know what is new say in a specific area and as some reader may not I often then write upon that in detail, for better or worse. I leave that up to the readers.
But sifting through masses of online data real time to see what has become new and how to fit that in to current research and writings is one of my organized ways. Twitter is totally useless. I have an account, I had to get one for something that I now forgot. I have Linkedin, and it has become the Ginza strip, all bright and useless. I left Facebook years ago, and Google+ makes no sense at all. There just seems to be stuff pasted by others and I have no idea why.
But if you are goal directed, with an organized mental bent, then Google Reader was spot on. If you are a Facebook fanatic, then it would make no sense. Google Reader was for those who did something with what they obtained, Facebook is a narcotic. It is Crazy Bird of the social set.
Google Reader led you to new ideas which made you modify and expand old ideas, it led to challenges, it led to thought. Google Reader was for the organized mind, not the idle mind.
Thus there is a potential wealth of information on how different people think. Social media is for those who want others to think for them. Perhaps a great marketing tool. Google Reader, plain and simple, is for those who want to think for themselves. This tells me something about the folks at Google. They are changing, and perhaps not for the best. Pity!
Labels:
Commentary,
Google
Friday, March 15, 2013
Beware The Ides of March
One should remember:
Soothsayer
Beware the ides of March.
Soothsayer
Caesar!CAESAR
Ha! who calls?CASCA
Bid every noise be still: peace yet again!CAESAR
Who is it in the press that calls on me? I hear a tongue, shriller than all the music, Cry 'Caesar!' Speak; Caesar is turn'd to hear.Soothsayer
Beware the ides of March.CAESAR
What man is that?BRUTUS
A soothsayer bids you beware the ides of March.CAESAR
Set him before me; let me see his face.CASSIUS
Fellow, come from the throng; look upon Caesar.CAESAR
What say'st thou to me now? speak once again.Soothsayer
Beware the ides of March.
Labels:
Commentary
Thursday, March 14, 2013
Now How Long Did This Take?
In 1967 I had designed a star tracking system used on the X 15 to managed updating the gyro for position location. We used a photomultiplier tube to scan and lock on to Polaris, the north star, and the earth's limb. It allowed precision position updates.
The in 1975 I added an optical, infrared, laser link between two Intelsat satellites as an option. We had designed the system and Bob Kennedy had delivered a detailed system design.
Now in 2013 I see the MIT Lincoln is finally adding an infrared link to a satellite to improve communications to earth from the satellite. Eureka states:
The LLCD mission will use a highly reliable infrared laser, similar to those used to bring high-speed data over fiber optic cables into our workplaces and homes. Data, sent in the form of hundreds of millions of short pulses of light every second, will be sent by the LADEE spacecraft to any one of three ground telescopes in New Mexico, California and Spain.
The real challenge of LLCD will be to point its very narrow laser beam accurately to ground stations across a distance of approximately 238,900 miles while moving. Failure to do so would cause a dropped signal or loss of communication.
"This pointing challenge is the equivalent of a golfer hitting a 'hole-in-one' from a distance of almost five miles," said Cornwell. "Developers at the Massachusetts Institute of Technology's (MIT) Lincoln Laboratory have designed a sophisticated system to cancel out the slightest spacecraft vibrations. This is in addition to dealing with other challenges of pointing and tracking the system from such a distance. We are excited about these advancements."
The LLCD mission will also serve as a pathfinder for the 2017 launch of NASA's Laser Communication Relay Demonstration (LCRD). That mission will demonstrate the long-term viability of laser communication from a geostationary relay satellite to Earth. In a geostationary orbit the spacecraft orbits at the same speed as Earth, which allows it to maintain the same position in the sky.
My only comments is that 40 years ago we demonstrated this, yes when I was also working at Lincoln. I just wonder what took so long.
The in 1975 I added an optical, infrared, laser link between two Intelsat satellites as an option. We had designed the system and Bob Kennedy had delivered a detailed system design.
Now in 2013 I see the MIT Lincoln is finally adding an infrared link to a satellite to improve communications to earth from the satellite. Eureka states:
The LLCD mission will use a highly reliable infrared laser, similar to those used to bring high-speed data over fiber optic cables into our workplaces and homes. Data, sent in the form of hundreds of millions of short pulses of light every second, will be sent by the LADEE spacecraft to any one of three ground telescopes in New Mexico, California and Spain.
The real challenge of LLCD will be to point its very narrow laser beam accurately to ground stations across a distance of approximately 238,900 miles while moving. Failure to do so would cause a dropped signal or loss of communication.
"This pointing challenge is the equivalent of a golfer hitting a 'hole-in-one' from a distance of almost five miles," said Cornwell. "Developers at the Massachusetts Institute of Technology's (MIT) Lincoln Laboratory have designed a sophisticated system to cancel out the slightest spacecraft vibrations. This is in addition to dealing with other challenges of pointing and tracking the system from such a distance. We are excited about these advancements."
The LLCD mission will also serve as a pathfinder for the 2017 launch of NASA's Laser Communication Relay Demonstration (LCRD). That mission will demonstrate the long-term viability of laser communication from a geostationary relay satellite to Earth. In a geostationary orbit the spacecraft orbits at the same speed as Earth, which allows it to maintain the same position in the sky.
My only comments is that 40 years ago we demonstrated this, yes when I was also working at Lincoln. I just wonder what took so long.
Labels:
NASA
Google, I have always had Questions
Now Google has a great search engine. Why, because unlike Yahoo it does not fill the screen up with junk. Do I make decisions to go to paid ad sites, yes quite often. But the best element of Google was, yes was, Google Reader. Well not any more.
You see the same morons who brought you Nexus 7, the Asus piece of junk, decided to cancel Google Reader. I use Google+ once a month at best. I am not a social media type. I left Facebook years ago, yes years ago, and I find Linkedin almost impossible to read, even on my 3 grand screens, on my gigantic desk. There is just too much noise.
But Google Reader has little if any noise. The BBC states:
A petition to save the service, which aggregates news content from web feeds, had 25,000 signatures in a few hours. Experts say shutting Reader is part of Google's plan to migrate more people to its social media service, Google+.
Who are these moron experts. I generally do not get this upset. It must be some media savvy twerp from Harvard. No one uses Google+, it stinks, so why close something that works to force people to something that does not. You see Silicon Valley is really filled with intellectual not wits. I bet they paid a fortune for this advice.
So what did I do, as soon as I got the news I ran for an alternative. You see I use Google Reader as a heads up for the latest research in my areas of interest. Get a hit, go to the paper, then write it up, add it to my ever increasing lore of stuff. My last two books would have been near impossible without Google Reader, I would even pay for it. At least until these neanderthals did this colossal misstep.
So what did I do? I went to Microsoft, yes the customer unfriendly Microsoft, and used that antediluvian piece of Microsoft code called Outlook, great little RSS unit. It took some tome, I used Google to get my feeds than loaded them into Outlook. A couple of hours of lost time working on my cancer models but necessary.
Now what did this tell me? First, cloud stuff really stinks. Outlook is not cloud, I have it on my system, thus some evil person cannot randomly remove it. Did I say evil, in reference to Google. Yes I did.
Second, Google seems to have gotten taken over by troglodytes. What happened folks. Are they going to stop the blogs as well, how about gmail, I don't use it anyhow, why not kill off the search engine too while you are at it.
This is clearly one of the stupidest actions taken by any company ever. Just look at the global backlash. Even in the NY Times, which states:
Now people will be out on the news reader street by July 1, and there are few places for them to go.
As BuzzFeed noted, using data from the BuzzFeed Network, a group of sites that collectively have over 300 million users,Google Reader still sends a considerable amount of traffic to these sites. Google Plus, the company’s social network, does not.
Remember, if all else fails, listen to the customer!
You see the same morons who brought you Nexus 7, the Asus piece of junk, decided to cancel Google Reader. I use Google+ once a month at best. I am not a social media type. I left Facebook years ago, yes years ago, and I find Linkedin almost impossible to read, even on my 3 grand screens, on my gigantic desk. There is just too much noise.
But Google Reader has little if any noise. The BBC states:
A petition to save the service, which aggregates news content from web feeds, had 25,000 signatures in a few hours. Experts say shutting Reader is part of Google's plan to migrate more people to its social media service, Google+.
Who are these moron experts. I generally do not get this upset. It must be some media savvy twerp from Harvard. No one uses Google+, it stinks, so why close something that works to force people to something that does not. You see Silicon Valley is really filled with intellectual not wits. I bet they paid a fortune for this advice.
So what did I do, as soon as I got the news I ran for an alternative. You see I use Google Reader as a heads up for the latest research in my areas of interest. Get a hit, go to the paper, then write it up, add it to my ever increasing lore of stuff. My last two books would have been near impossible without Google Reader, I would even pay for it. At least until these neanderthals did this colossal misstep.
So what did I do? I went to Microsoft, yes the customer unfriendly Microsoft, and used that antediluvian piece of Microsoft code called Outlook, great little RSS unit. It took some tome, I used Google to get my feeds than loaded them into Outlook. A couple of hours of lost time working on my cancer models but necessary.
Now what did this tell me? First, cloud stuff really stinks. Outlook is not cloud, I have it on my system, thus some evil person cannot randomly remove it. Did I say evil, in reference to Google. Yes I did.
Second, Google seems to have gotten taken over by troglodytes. What happened folks. Are they going to stop the blogs as well, how about gmail, I don't use it anyhow, why not kill off the search engine too while you are at it.
This is clearly one of the stupidest actions taken by any company ever. Just look at the global backlash. Even in the NY Times, which states:
Now people will be out on the news reader street by July 1, and there are few places for them to go.
As BuzzFeed noted, using data from the BuzzFeed Network, a group of sites that collectively have over 300 million users,Google Reader still sends a considerable amount of traffic to these sites. Google Plus, the company’s social network, does not.
Remember, if all else fails, listen to the customer!
Labels:
Google
Sunday, March 10, 2013
Employment: Getting Better?
First the Romer Curve. No we have not yet gotten to where she said.
Yes her projections are still way off. And yes she seems to be indoctrinating more minds as to why this has not been the case.
The good news is increasing participation. Notice the upswing as consistent.
If I believe the numbers this actually looks good. We seem to be going back up.
The above reinforces that. Hopefully this is sustainable.
Yes her projections are still way off. And yes she seems to be indoctrinating more minds as to why this has not been the case.
The good news is increasing participation. Notice the upswing as consistent.
If I believe the numbers this actually looks good. We seem to be going back up.
The above reinforces that. Hopefully this is sustainable.
Labels:
Economy
Saturday, March 9, 2013
Banting and Best, What is the Cause Again?
Banting and Best where are you when we really need you? I am
always amazed by authors such as Basu et al who come out with a new discovery
regarding sugar and Diabetes. It is as if they have just gotten some clear
insight into the obvious. Obesity drives the suppression of the islet cells
which drives down the insulin production which drives up the glucose level. Get
the BMI below 22.5 and you drive out the main initiator of Type 2 Diabetes, the
inflammatory effects of weight.
Now the authors state[1]:
While experimental and observational studies suggest that
sugar intake is associated with the development of type 2 diabetes, independent
of its role in obesity, it is unclear whether alterations in sugar intake can
account for differences in diabetes prevalence among overall populations.
Using econometric models of repeated cross-sectional data
on diabetes and nutritional components of food from 175 countries, we found
that every 150 kcal/person/day increase in sugar availability (about one can of
soda/day) was associated with increased diabetes prevalence by 1.1% (p ,0.001)
after testing for potential selection biases and controlling for other food
types (including fibers, meats, fruits, oils, cereals), total calories, overweight
and obesity, period-effects, and several socioeconomic variables such as aging,
urbanization and income.
No other food types yielded significant individual
associations with diabetes prevalence after controlling for obesity and other confounders.
The impact of sugar on diabetes was independent of sedentary behavior and
alcohol use, and the effect was modified but not confounded by obesity or
overweight.
Duration and degree of sugar exposure correlated
significantly with diabetes prevalence in a dose-dependent manner, while
declines in sugar exposure correlated with significant subsequent declines in
diabetes rates independently of other socioeconomic, dietary and obesity
prevalence changes. …
In summary, population-level variations in diabetes
prevalence that are unexplained by other common variables appear to be statistically
explained by sugar. This finding lends credence to the notion that further
investigations into sugar availability and/or consumption are warranted to
further elucidate the pathogenesis of diabetes at an individual level and the
drivers of diabetes at a population level.
Well one would think that to be the case. Sugar, sucrose, is
a powerful carbohydrate and Diabetes is a carbohydrate disorder, and Type II
Diabetes is exacerbated by obesity and sugar has excess calories thus causing
obesity. It is logical causality. We have seen genes, billboards, environment,
sugar, and endless excuses for the fact that people eat too much.
Most physicians treating Type II Diabetes sees the patient
as obese, having a high carb diet, more than likely smoking and with no
exercise. Further if they examine the diet, they find tremendous amounts of
carbs, and if they look deeper they are dominated by various sugars. Thus this
conclusion is not only obvious but it faces physicians on a day by day basis.
The prototypical Type II Diabetic is a candy eating obese
person. Instead of a small candy bar, they get bags of them, cakes, cookies,
and the list continues. So what is new?
Labels:
Obesity
Friday, March 8, 2013
The Cause of Obesity: It is Food Stupid!
There is an article in today's NY Times which states:
OBESITY is a problem everywhere, with significant consequences for personal health and public spending. People weigh more than ever — but why? If we can find the causes of obesity, we can try to eliminate or counter them. Unfortunately, finding causes is easier said than done, and causes we think we see can turn out to be illusions.
The cause, well simple, it is food. Remember 3500 kcal per pound. You basal met rate is say 2000 kcal per day, you eat 2500 kcal per day you gain a pound a week. Simple, End of story. Do not look any further! Input less output is Net Accumulation.
Now here is a real dumb statement:
The gold standard for inferring causation in social science, as in medicine, is the randomized controlled trial, in which people or places are randomly assigned to receive different treatments. In this case, the “treatment” would be the amount of outdoor food ads in an area. But advertisers are unlikely to agree to randomly distribute their signs, nor would people consent to live in a randomly chosen place.
The gold standard is simple physics, in this case energy input and output. The author is a psychologist so I guess I should not be surprised. It is food, too much food. Let me give a few examples:
1. A decade ago when I brought my Czech partners and their family to the US I took them to a lunch restaurant. The plates came out and they were shocked, they at first thought one plate was for all, like a family meal, then they realized that one was for each. In Prague at lunch we had a small plate of food, and even 6 oz of beer, not my choice, but all in moderation.
2. At a day recently of lunches and dinners I kept to my bowl of soup while I saw others with fries and piles of food, well in excess of the 2000 kcal per day. It was not pure choice, but it was choosing Item 9 on the menu. I went off menu. No problem, just ask.
3. The typical business lunch in a conference room contains sandwiches that contain not only bread and meat but a collection of calorie adding stuff. The solution, remove the bread and stuff. Stick with the protein. Also there is always the soda and deserts. Hands off!
The "cause" of obesity is obvious, more input than output. The more I speak with people the more I am concerned the only way to stop it is through taxation, pure and simple. Especially if those who opine like those in the Times are totally clueless.
OBESITY is a problem everywhere, with significant consequences for personal health and public spending. People weigh more than ever — but why? If we can find the causes of obesity, we can try to eliminate or counter them. Unfortunately, finding causes is easier said than done, and causes we think we see can turn out to be illusions.
The cause, well simple, it is food. Remember 3500 kcal per pound. You basal met rate is say 2000 kcal per day, you eat 2500 kcal per day you gain a pound a week. Simple, End of story. Do not look any further! Input less output is Net Accumulation.
Now here is a real dumb statement:
The gold standard for inferring causation in social science, as in medicine, is the randomized controlled trial, in which people or places are randomly assigned to receive different treatments. In this case, the “treatment” would be the amount of outdoor food ads in an area. But advertisers are unlikely to agree to randomly distribute their signs, nor would people consent to live in a randomly chosen place.
The gold standard is simple physics, in this case energy input and output. The author is a psychologist so I guess I should not be surprised. It is food, too much food. Let me give a few examples:
1. A decade ago when I brought my Czech partners and their family to the US I took them to a lunch restaurant. The plates came out and they were shocked, they at first thought one plate was for all, like a family meal, then they realized that one was for each. In Prague at lunch we had a small plate of food, and even 6 oz of beer, not my choice, but all in moderation.
2. At a day recently of lunches and dinners I kept to my bowl of soup while I saw others with fries and piles of food, well in excess of the 2000 kcal per day. It was not pure choice, but it was choosing Item 9 on the menu. I went off menu. No problem, just ask.
3. The typical business lunch in a conference room contains sandwiches that contain not only bread and meat but a collection of calorie adding stuff. The solution, remove the bread and stuff. Stick with the protein. Also there is always the soda and deserts. Hands off!
The "cause" of obesity is obvious, more input than output. The more I speak with people the more I am concerned the only way to stop it is through taxation, pure and simple. Especially if those who opine like those in the Times are totally clueless.
Labels:
Obesity
Thursday, March 7, 2013
Age of Edison: Review
The Age of Edison by Freeberg is a compelling tale of technological
innovation and the machinations and creations of all those who participated. At
the center is Edison, whose fame was a creation of what he accomplished, what
he proclaimed, and what the Press found as good news copy, independent of the
reality of what was truly happening. Mr Freeberg has written an exceptionally well balanced and factual book. Unlike the book, The Idea Factory, about Bell Labs, Freeberg looks at each player evenly and based upon facts. His view is expansive and he does not appear to make saints out of sinners. Thus unlike The Idea Factory which is replete with positive statements where the truth may actually be different, Freeberg presents the facts and the result is masterful.
I live but a short distance from Edison’s last lab was in West Orange in New Jersey and it is now a National Parks site. Much of what Edison did is memorialized by the many labs, books, and remnants of his hundreds of “inventions”. Of course next to this National Landmark is the Edison battery factory which one may suppose is left in a state of total collapse because the cleanup of the site would be astronomical, but those factors are somewhat missing from the tale.
I live but a short distance from Edison’s last lab was in West Orange in New Jersey and it is now a National Parks site. Much of what Edison did is memorialized by the many labs, books, and remnants of his hundreds of “inventions”. Of course next to this National Landmark is the Edison battery factory which one may suppose is left in a state of total collapse because the cleanup of the site would be astronomical, but those factors are somewhat missing from the tale.
The book is exceptionally well written and it is really a
tale of the electric light, with Edison cast as someone who comes and goes, and
yet has a lasting influence. Like so many technological advances there is usually
not one person, but many competing for the prize. The goal was clear, light,
but the path uncertain. The author details the competition between the arc
light and the incandescent light, the need for an infrastructure, and the
problems of that infrastructure. Power lines grew, collided with humanity, and
in urban areas were driven underground. However they remained to be smashed
down during hurricane Sandy, almost 150 years after all of this began. Thus the
power industry, unlike the electronics industry rapidly grew, and then froze,
for almost a century. But this is a tale of the light bulb, perhaps the most
significant driver of that industry.
The author opens with the inventing of the light bulb. He
wonderfully shows with balance and insight many if not most of the players
during this time. Edison was thus one of many, but perhaps the most effective
self-promoter. He also had strong financial backers who used their strength as
well.
The author then discusses the diffusion of light to both
work environments as well as leisure environments. He does a great job showing
hos this diffusion changed the way people interacted. This is a critical
observation of how technology effects sociological change.
The author discusses the whole issue of patents and patent
battles. At the time of Edison there was a strong development of Socialist
movements in the US. On p 153 the author discusses the battles over patents.
Socialists as he says:
“People invented to satisfy natural creative urge, the socialists
insisted, and out of desire to help others. But capitalists bought up the
patent rights.”
In a sense the author describes the same battle we see today
with some on the Internet who feel that content should be free, and that
copyright rights are to be trampled.
The author discusses the expansion of applications, some
good and some useless. On p 169 he describes certain medical applications, some
good some useless. Yet at the same time we see the invention of the X-ray
systems, which in a way was a natural step from the incandescent light bulb.
The battle between “standards” is also brought out by the
author in the battle between AC and DC. In reading of Edison’s views, for he
was a DC promoter due to his collections of patents in that space, he never did
grasp the basic truth that high voltage AC, using transformers, allowed for
very low loss transmission over long distances. Edison apparently just did not
understand the theory, unlike Tesla, who was a well-educated engineer. Edison
was a technician at best, and when that failed he had a large collection of
technicians, but in reviewing his library he had little along the lines of true
technology. He had technique, a technique developed by extensive trial and
error.
On p 199 the author discusses the issue of municipal
ownership of utilities. Specifically he talks of the strong Progressive drive
to have municipalities control such vital resources. In fact they wanted to
control telephone and telegraph, water and sewer. Again what the author has done
is to lay out the issue as the technology evolved and he demonstrates so well
the mapping on today’s same issues in such areas as broadband. In a sense this
book uses the light bulb to demonstrate a near universal development process,
sociologically and politically, of almost any new massively accepted
technology.
On p 205 the author recounts the development of the
technologists, the introduction of electrical engineering into universities
such as MIT, Cornell, and Columbia. In this case the universities were
followers; they were presented with a pile of technology driven by techniques
with no well accepted basis for growth, and then began constructing the basis.
Later in the book the author returns to Government control
over the diffusion of this technology. On p 301 is a discussion of the New Deal
and the Rural Electrification Administration, bringing light to the farmer. As
he says:
“The New Deal’s social engineers believed that rural
electrification would do much to ease the burden of farm work…”
Also he notes the FDR administration wanted to bridge what
they saw as a growing gap of rural and urban America. Again the author has
brilliantly carried the tale to an end point and a point which we can see again
today in the broadband arguments.
Overall this book serves two purposes. First it is an
excellent summary of the evolution of the light bulb across many facets of
society. Second and I believe more important, it represents a paradigm for
understanding the development, diffusion and politicization of technological
change.
Labels:
Books,
Technology
Subscribe to:
Posts (Atom)